Nucleic Acids Research

The authors wish it to be known that, in their opinion, the first five authors should be regarded as Joint First Authors.
- Altmetric
Many studies have indicated that non-coding RNA (ncRNA) dysfunction is closely related to numerous diseases. Recently, accumulated ncRNA–disease associations have made related databases insufficient to meet the demands of biomedical research. The constant updating of ncRNA–disease resources has become essential. Here, we have updated the mammal ncRNA–disease repository (MNDR, http://www.rna-society.org/mndr/) to version 3.0, containing more than one million entries, four-fold increment in data compared to the previous version. Experimental and predicted circRNA–disease associations have been integrated, increasing the number of categories of ncRNAs to five, and the number of mammalian species to 11. Moreover, ncRNA–disease related drug annotations and associations, as well as ncRNA subcellular localizations and interactions, were added. In addition, three ncRNA–disease (miRNA/lncRNA/circRNA) prediction tools were provided, and the website was also optimized, making it more practical and user-friendly. In summary, MNDR v3.0 will be a valuable resource for the investigation of disease mechanisms and clinical treatment strategies.
INTRODUCTION
The associations between ncRNA dysfunction and diseases have been the focus of attention in recent decades (1–5). With the continuous advancement of sequencing technology and prediction algorithms, experimentally validated and computationally predicted ncRNA–disease associations have explosively increased. Some ncRNAs, such as circRNAs whose functions were once unclear, have also been found to be closely related to diseases recently (6–10). The increasing growth of data requires that the existing ncRNA–disease data resources must be constantly updated to satisfy the requirements of disease research and clinical applications, such as, to our best knowledge, masses of piRNAs were found dysregulated in Parkinson's disease (11), but not collected in any related databases. In addition, the research on ncRNA and drugs has also been developed rapidly. For example, some long non-coding RNAs (lncRNAs) in cancer may present potential therapeutic targets, and both microRNA (miRNA) and lncRNA have been reported to play important roles in drug resistance (12,13). However the collection and integration of related drugs are still insufficient, which limits the research on the association and mechanism between drugs, ncRNA and diseases. Furthermore, some studies have indicated that the subcellular localization and interaction of ncRNA could also affect diseases (14,15). Accordingly, it is essential to update the relevant database in real time.
Because of the above factors, this version of the mammal ncRNA–disease repository (MNDR, http://www.rna-society.org/mndr/) was brought into being. We integrated different kinds of ncRNA–disease associations through manual literature curation and prediction algorithms, with other resources under one common framework. Compared to the pervious release, the update mainly improves the following aspects: (i) more than one million entries, four-fold increment in data, and an increase to 11 mammals; (ii) the addition of circRNA associations; (iii) the addition of drug-related information; (iv) the integration of ncRNA subcellular localization and interaction, (v) support for three ncRNA–disease prediction tools and (vi) more user-friendly interface and web services were designed. In summary, MNDR v3.0 provides comprehensive data on ncRNA–disease associations in mammals, helping to better understand the mechanism of ncRNAs and diseases.
DATA COLLECTION AND ORGNIZATION
MNDR v3.0 contains experimentally validated and computationally predicted ncRNA–disease associations from the literature and other resources, respectively. We have reviewed over 25 000 published studies and acquired >40 000 experimental ncRNA–disease associations. The diverse ncRNA–disease associations from 17 related experimentally validate databases (16–32) and 14 computationally predicted algorithms (31–44) were also integrated (Supplemental Table S1). In extension, drug-related information was obtained from four databases: ncDR (45), NoncoRNA (46), NRDTD (47) and RNAInter (48), ncRNA subcellular localizations were obtained from RNALocate (15) and interactions from RNAInter (48).
To unifying the data from different sources into authoritative reference databases, lncRNA symbols were mapping to the NCBI gene and Ensembl (49), while miRNA, circRNA and piRNA symbols to miRbase (50), circBase (51) and piRBase (52), respectively. Sno/scaRNAbase (53) and snoRNA-LBME-db (54) were chosen for snoRNA symbols. The disease terms were mapping to the Disease Ontology (55) and MeSH vocabularies. Related drug annotations were selected from PubChem Compound.
RESULTS
MNDR v3.0 statistics
In total, MNDR v3.0 contains 1 007 831 ncRNA–disease associations, across 11 mammals and documents 24 323 publications (Figure 1). Regarding prediction data, MNDR v3.0 includes 237 329 miRNA-associated, 252 144 lncRNA-associated and 296 910 circRNA-associated entries for Homo sapiens, as well as 2434 and 28 predicted lncRNA–disease associations for Mus musculus and Rattus norvegicus, respectively. Compared to the pervious release, number of the species was increased from 6 to 11 in MNDR v3.0 (Table 1). There are a total of 6301 non-redundant miRNAs, together with 39 880 lncRNAs, 20 506 circRNAs, 10 894 piRNAs and 521 snoRNAs, and the number of types of diseases was increased to 1614. In addition, related drug annotations and four types of ncRNA-drug associations: drug target, drug sensitive, drug resistant and drug interaction were also included.

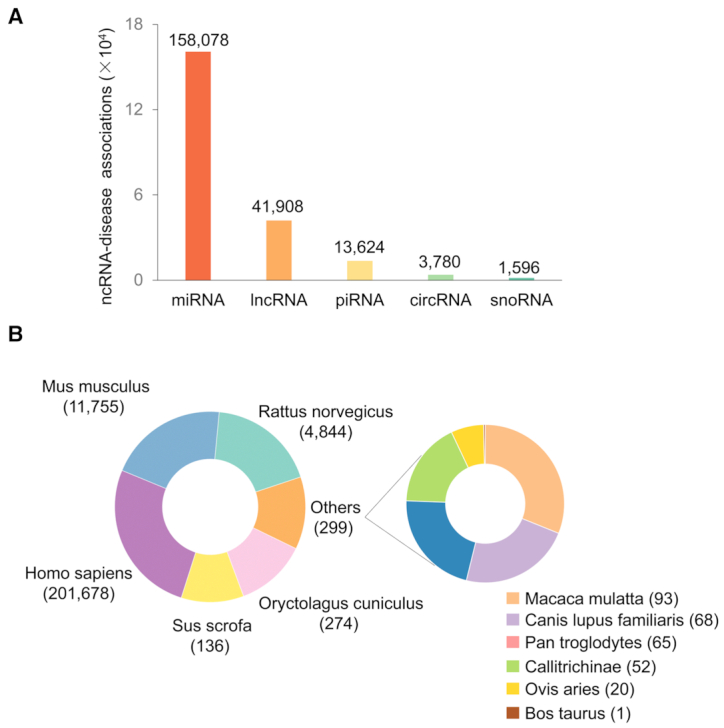
Statistics on MNDR v3.0. (A) The distribution of experimental ncRNA–disease associations in five types of ncRNA (miRNA/lncRNA/circRNA/piRNA/snoRNA). (B) Number of experimental associations in 11 mammals.

| Feature | MNDR v1.0 | MNDR v2.0 | MNDR v3.0 |
|---|---|---|---|
| Entry | 1149 | 261042 | 1007831 |
| RNA symbol | 369 | 23858 | 78102 |
| Disease | 175 | 1416 | 1614 |
| Specie | 3 | 6 | 11 |
| Literature | 377 | 11504 | 24323 |
| RNA category | miRNA/lncRNA/piRNA | miRNA/lncRNA/piRNA | miRNA/lncRNA/circRNA |
| /snoRNA | /snoRNA | /piRNA/snoRNA | |
| Detailed information | Basic annotation | Basic annotation | Basic annotation |
| Evidence support | DO/MeSH description | DO/MeSH description | |
| Reference | Evidence support | Drug information | |
| Reference | RNA interaction | ||
| RNA localization | |||
| Evidence support | |||
| Reference | |||
| Web application | - | Browse | Browse |
| Advanced filter search | Exact search/Fuzzy search | ||
| /Batch search | |||
| Three prediction tools: | |||
| SPM, SIMCLDA, DeepDCR |
Database usage
To satisfy the different requests of biomedical researchers, a more user-friendly web interface and convenient search and browse functions have been designed in MNDR v3.0. It enables an optimized query with new fuzzy and batch functions. Users can use ‘Fuzzy Search’ to search ncRNA–disease associations by unstandardized or uncertain ncRNA name/disease name and then choose further from the candidate list. ‘Batch Search’ supports inputting a list of ncRNA official symbols/IDs, and disease names/IDs (DOID/MeSH ID), as well as uploading a file in text format to obtain multiple ncRNA–disease associations. By doing so, users can select ‘Exact Search’ to filter the search results, ‘Fuzzy Search’ to further focus on ncRNA or disease of interest, or ‘Batch Search’ to customize their query content by batch. The search results can be downloaded by clicking the button above the result table. MNDR v3.0 also offers a download option on the ‘Browse’ page.
Prediction tools
MNDR v3.0 provides three ncRNA–disease prediction tools on the website (Figure 2): SPM (structural perturbation method) was used for miRNA–disease prediction (56), while SIMCLDA based on inductive matrix completion was applied for the lncRNA–disease prediction (57), and the deep forests joint positive-unlabeled learning algorithm DeepDCR could be used to calculate the associations between circRNAs and diseases (42).
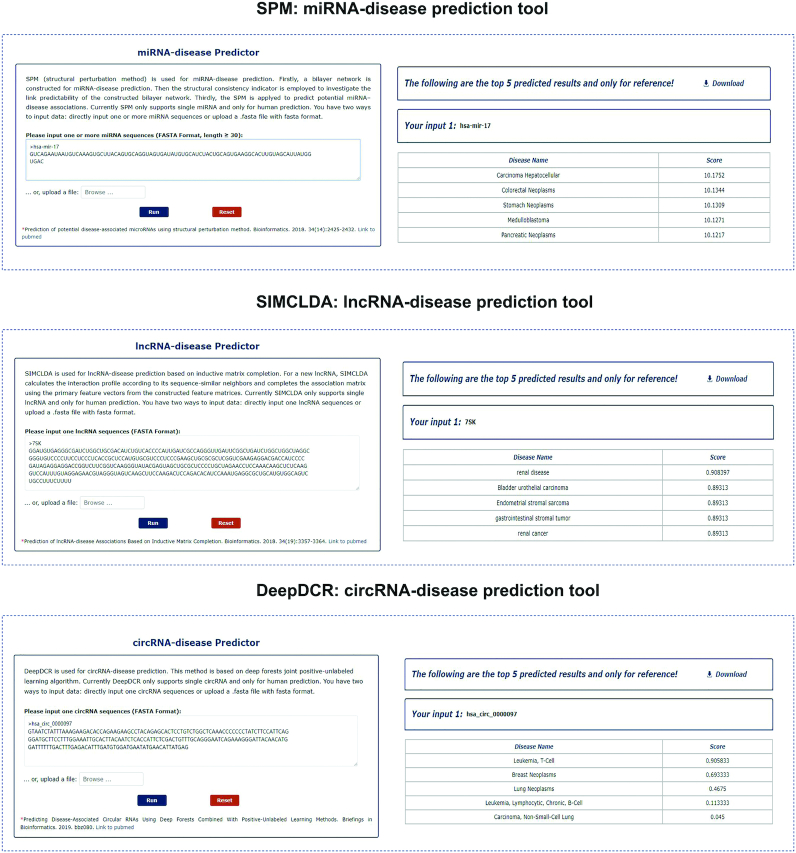


Snapshot of three ncRNA–disease prediction tools in MNDR: SPM, SIMCLDA and DeepDCR (left: input option, right: the presentation of results).
CONCLUSIONS AND PERSPECTIVES
With the continuous development of high throughout technologies and predictive algorithms, the evidence of ncRNA–disease associations has increased greatly in the recent years. Meanwhile, research on the ternary relationships between related drugs, ncRNAs and diseases is receiving increasing attention, and ncRNA subcellular localizations and interactions are also confirmed to be related with the regulation of diseases. To address the above aspects, the MNDR database was updated with the latest data and some new and improved features. MNDR v3.0 contains over one million entries, including five types of ncRNA, covering 11 mammals. With the massive growth of associations, the diversification of annotations and the optimization of website interface and functions, MNDR v3.0 depicts a system-level ncRNA–disease landscape, helping researchers obtain accurate and comprehensive data more conveniently for further exploration. We may optimize the evaluation algorithm to respond to accumulating ncRNA–disease associations in the future, and will continually maintain and update MNDR database to satisfy the growing requirements for the investigation of disease mechanisms and related clinical applications.
SUPPLEMENTARY DATA
Supplementary Data are available at NAR Online.
FUNDING
National Key Research and Development Project of China [2019YFA0801800]; National Natural Science Foundation of China [81770104]; Basic and Applied Basic Research Fund of Guangdong Province [2019A1515010784, 2019A1515110701]. Funding for open access charge: National Key Research and Development Project of China [2019YFA0801800]; National Natural Science Foundation of China [81770104]; Basic and Applied Basic Research Fund of Guangdong Province [2019A1515010784, 2019A1515110701].
Conflict of interest statement. None declared.
REFERENCES
3.
4.
5.
6.
7.
8.
9.
10.
11.
12.
13.
14.
15.
16.
17.
18.
19.
20.
21.
22.
23.
24.
25.
26.
27.
28.
29.
30.
31.
32.
33.
34.
35.
36.
37.
38.
39.
40.
41.
42.
43.
44.
45.
46.
47.
48.
49.
50.
51.
52.
53.
54.
55.
56.
 MNDR v3.0: mammal ncRNA–disease repository with increased coverage and annotation
MNDR v3.0: mammal ncRNA–disease repository with increased coverage and annotation
